Spacia: Inferring Cell-cell Interaction from Spatial Transcriptomics Data
Multicellular organisms heavily rely on cell-cell interactions to effectively coordinate and regulate various biological processes, ensuring the normal functioning of the organism. Spacia models and evaluates cell-cell interactions from spatial transcriptomic data (SRT). This model uses cell-cell proximity as a constraint and prioritizes cell-cell interactions that cause a downstream change in the cells. Spacia employs a Bayesian multi-instance learning (MIL) framework to assess intercellular communication from between cells and their neighbors.
Graphical abstract
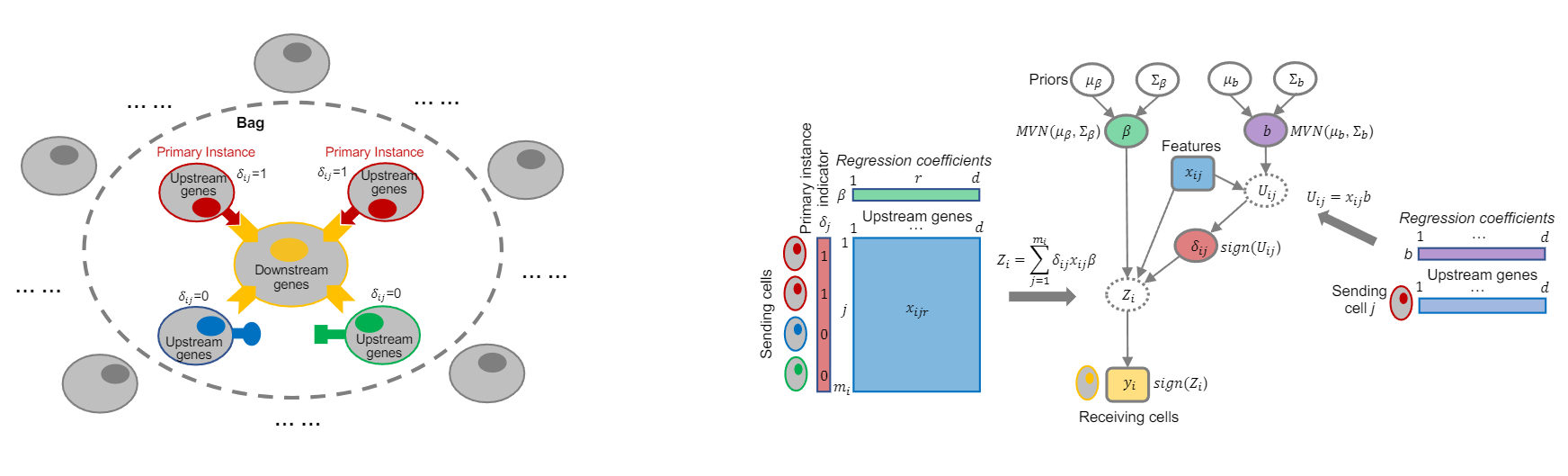
Installation
Usage
Contact Us
If you have any suggestions/ideas for Spacia or are having issues trying to use it, please don’t hesitate to reach out to us.
Noah Chang, wooyong.chang@utsouthwestern.edu
James Zhu, james.zhu@utsouthwestern.edu
Dr. Yunguan Wang, yunguan.wang@utsouthestern.edu
For more resources on our labs, collaborators, and related projects, please visit the following:
For project development and newest updates, consider visiting the following:
Our Github